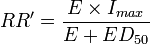Exposure-response function
| Moderator:Jouni (see all) |
|
|
| Upload data
|
- Exposure-response function (ERF) (or exposure-response relationship) is the relationship between the exposure of a given organism, system, or (sub)population to an agent in a specific pattern during a given time and the magnitude of a continuously graded effect to that organism, system, or (sub)population.
- This term has several related terms that may have slightly different meaning. Effect and response are interchangeable words. Also the word function is used instead of relationship. In Opasnet, we use the term exposure-response function (or ERF) as the generic term for different kinds of relationships. Often the exposure metric is more specifically defined in an alternative term. Two common examples:
- Concentration-effect relationship
- Relationship between the exposure, expressed in concentration, of a given organism, system, or (sub)population to an agent in a specific pattern during a given time and the magnitude of a continuously graded effect to that organism, system, or (sub)population. The concentration is measured at a defined site. [1]
- Dose-response relationship
- Relationship between the amount of an agent administered to, taken up by, or absorbed by an organism, system, or (sub)population and the change developed in that organism, system,or (sub)population in reaction to the agent. [1]<section end=glossary />
Contents
Question
What is such a representation for ERF that it fulfills the following criteria?
- It is widely applicable to all kinds of agents, exposures, and responses.
- A single ERF is widely applicable, within its domain, to different situations and populations.
- It is mathematically clear so that impact calculations can be operationalised based on it.
Answer
An ERF is a mathematical construct describing the relationship between a response and an exposure. In the general form, it is described as a probability.
Rationale
Equations
By nature, a response to an agent occurs at individual level (with some exceptions such as agents affecting herd immunity). Thus, the individual response should be the basic unit for ERF. If individual variation is of no interest, a population ERF can simply be expressed as the average of individual ERFs.
There are several different functions that may be used. These are defined here. In all equations, these variables are used:
- RR: the relative risk for unit exposure difference. Note that β = ln(RR).
- E: exposure level of the individual
- B: background exposure level that is considered negligible or lowest achievable. Exposures below B are not considered.
- T: threshold exposure level below which no impact will occur
- RR' is the relative risk for the actual exposure
- Imax is the maximal relative impact
- ED50 is the dose that causes 50 % of the maximal impact.
NOTE! ED50 parameter is given in the Threshold column, and Imax parameter is given in the ERF or Result column.
Approaches relative to background disease risk
- Relative risk (RR)
- describes the relative risk compared with a reference exposure. The actual number of cases is calculated with the equation below. (E > T) is 1 if E is greater than T and 0 otherwise.

- Relative Hill
- Relative Hill means an ERF function derived from Hill's plot:
Approaches independent of disease risk
- Exposure-response slope (ERS)
- A linear relationship where ERS defines the slope of the exposure-response line. Typically the intercept is assumed to be 0. Response metric MUST be defined as it varies from one case to another.

- Cancer slope factor (CSF)
- A linear relationship between constant lifetime exposure (typically in units mg/kg/d) and lifetime probability of cancer.

- Step function
- Assumes that exposure at certain range is not a hazard, while exposure outside that range is. Threshold and ERF parameters are used to give the lower and upper ends of the range, respectively. The Step function includes tolerable daily intake (TDI), recommended daily intake (RDI), acceptable daily intake (ADI), no-observed-adverse-effect level (NOAEL) and related, often administrative limits. The value is TRUE (or 1) if exposure fails to meet the recommendation and FALSE (or 0) otherwise.
Pages with standardised ERF tables
If a page has a standardised ERF data table (see an example above), the data can be automatically read and used by an R code and combined with other ERF tables. Therefore it is preferable to use the standard format. It makes modelling much easier and also enables an easy way to add more endpoints to assessments, if there are interesting exposures and available ERF tables.
The tables can contain the following columns (obligatory columns are in bold):
- Obs (automatic)
- Exposure agent (index)
- Response (index)
- Population (index)
- Age (index)
- Sex (index)
- Exposure (index)
- Exposure unit (index)
- ER function (index)
- Exposure metric (index)
- Scaling (index)
- Observation (hidden), containing two locations:
- Threshold
- ERF
- Result
- Description (description, there may be any number of description columns because they are not stored in the database)
Note! Spaces in column names will be replaced with "_" to avoid problems in the code.
| Page | Ident | Code name | Description |
|---|---|---|---|
| ERFs of environmental pollutants | Op_en5827 | ERF_env2 | Contains ERFs for radon, PM2.5, noise, chlorinated byproducts in drinking water, arsenic, dampness in buildings, formaldehyde, fluoride, ozone, lead, dioxin, quartz dust, asbestos. |
| ERF of omega-3 fatty acids | Op_en5830 | ERF_omega32 | Contains ERFs for Omega3 fatty acids. |
| ERF of methylmercury | Op_en5825 | ERF_mehg2 | Contains ERFs for MeHg. |
| ERF of dioxin | Op_en5823 | ERF_diox2 | Contains ERFs for dioxin TEQ. |
| ERFs of vitamins | Op_en6866 | ERF_vit2 | Contains ERF for vitamin D. |
| ERF of waterborne microbes | Op_en7948 | ERF_micr2 | Contains ERF for vitamin D. |
| ERF of PFAS | Op_en7974 | ERF_pfas | Contains ERF for perfluorinated alkyl substances |
Calculations
Technical issues
- In Opasnet, the use of term Exposure-response function (ERF) is recommended. The pages about ERFs should be named: "ERF of <agent> on <endpoint> in <population>."
- ERFs are typically variables, and they should be categorised to Category:Exposure-response functions
- In the definition of a variable, an ERF data is described as a t2b table under subheading Data. The actual ovariable that is used in models is defined in an "initiate" code under subheading Calculations.
See also
- heande:File:2013 04 03- Exposure-Response Functions DRAFT.xlsx ERFs for Urgenche
- A set of exposure-response functions from NEEDS project.
- Description of the ERF of Lden on annoyance, to exemplify the use of an ERF with a % affected type of response metric (rather than the more common "relative risk").
References
Weihsueh A. Chiu and Wout Slob. (2015) A Unified Probabilistic Framework for Dose–Response Assessment of Human Health Effects. Environ Health Perspect; doi:10.1289/ehp.1409385